|
Secondly, It can be used in generation of sequence/structure alignment. Firstly, dihedral angle prediction may act as substitute or supplement for secondary structure prediction. On the whole, dihedral angle prediction may benefit protein tertiary structure prediction in several aspects. Thus, prediction of dihedral angles is especially useful for protein tertiary structure prediction. The local structural bias information restricts the possible conformations of a sequence segment and therefore narrows down the conformation space of the whole polypeptide chain significantly. It is a significant step towards establishing the structure and function of a protein to predict local conformation of the polypeptide chain. However, secondary structure states are described as discrete classes and there is no clear boundary between coil and helical/strand states. It has been shown that predicted secondary structure is useful in the prediction of disordered and flexible regions, fold recognition and function prediction. reviewed the progress in the field of intermediate state or one-dimensional property prediction. That means to transform the ultimate goal into several sub-problems, such as secondary structure prediction, solvent accessibility prediction, residue-residue contact prediction, etc. But it is challenging to directly predict tertiary structure from primary sequence, so the hierarchical approach has been widely accepted as one of the most efficient methods. It has been shown that sequences contain rich information for protein tertiary structure prediction as well as functional study. Our study provides an alternative and more accurate prediction of dihedral angles, which may facilitate protein structure prediction and functional study. That is, the real prediction error can be well approximated by our estimated bounds. Our result also shows approximately linear relationship between the real prediction errors and our estimated bounds. 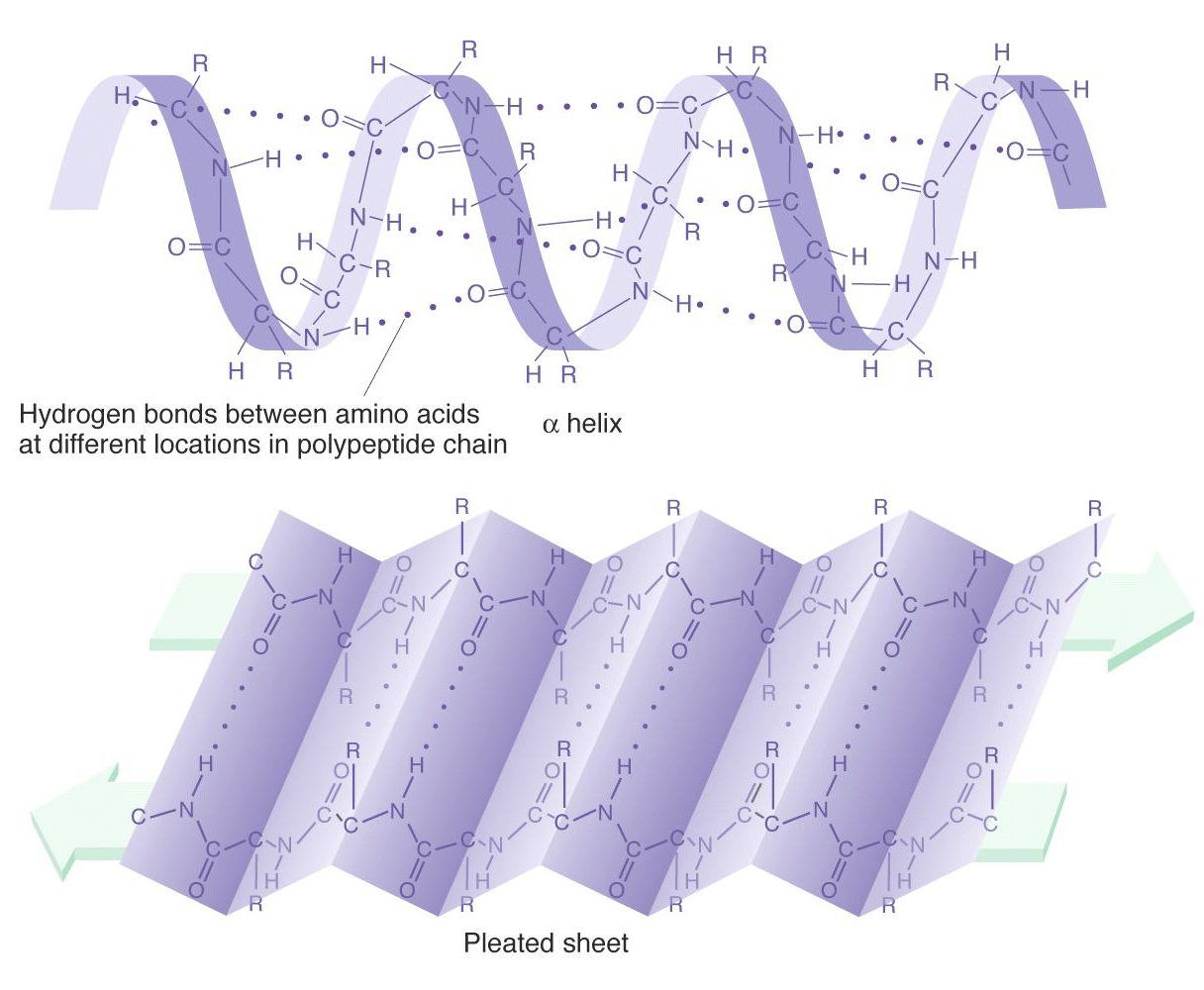
Tested on a subset of PDB25 and the targets in the latest two Critical Assessment of protein Structure Prediction (CASP), our method outperforms the existing state-of-art method SPIDER2 in terms of Pearson Correlation Coefficient (PCC) and Mean Absolute Error (MAE). In this article, we present a novel method (named RaptorX-Angle) to predict real-valued angles by combining clustering and deep learning. However, direct angle prediction from sequence alone is challenging. 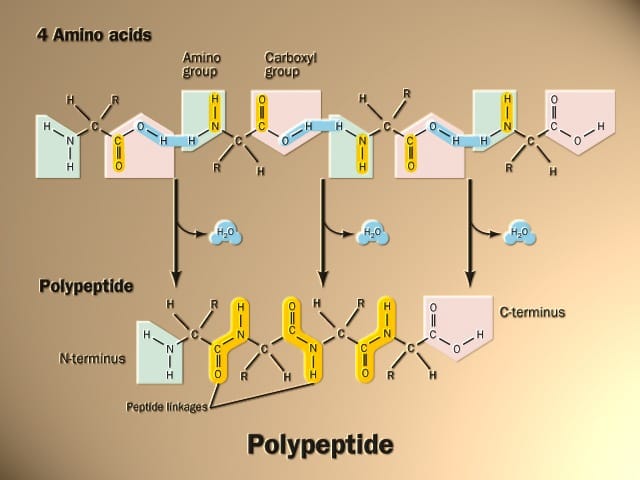
Predicted dihedral angles can be used to narrow down the conformational space of the whole polypeptide chain significantly, thus aiding protein tertiary structure prediction. Protein dihedral angles provide a detailed description of protein local conformation.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
